Base R Data Structures: Vectors
Common Data Structures
A data scientist needs to deal with data! We need to have a firm foundation in the ways that we can store our data in R. This section goes through the most commonly used ‘built-in’ R objects that we’ll use.
There are five major data structures used in
R- Atomic Vector (1d)
- Matrix (2d)
- Array (nd)
- Data Frame (2d)
- List (1d)
| Dimension | Homogeneous (elements all the same) | Heterogeneous (elements may differ) |
|---|---|---|
| 1d | Atomic Vector | List |
| 2d | Matrix | Data Frame |
| nd | Array |
(Atomic) Vector
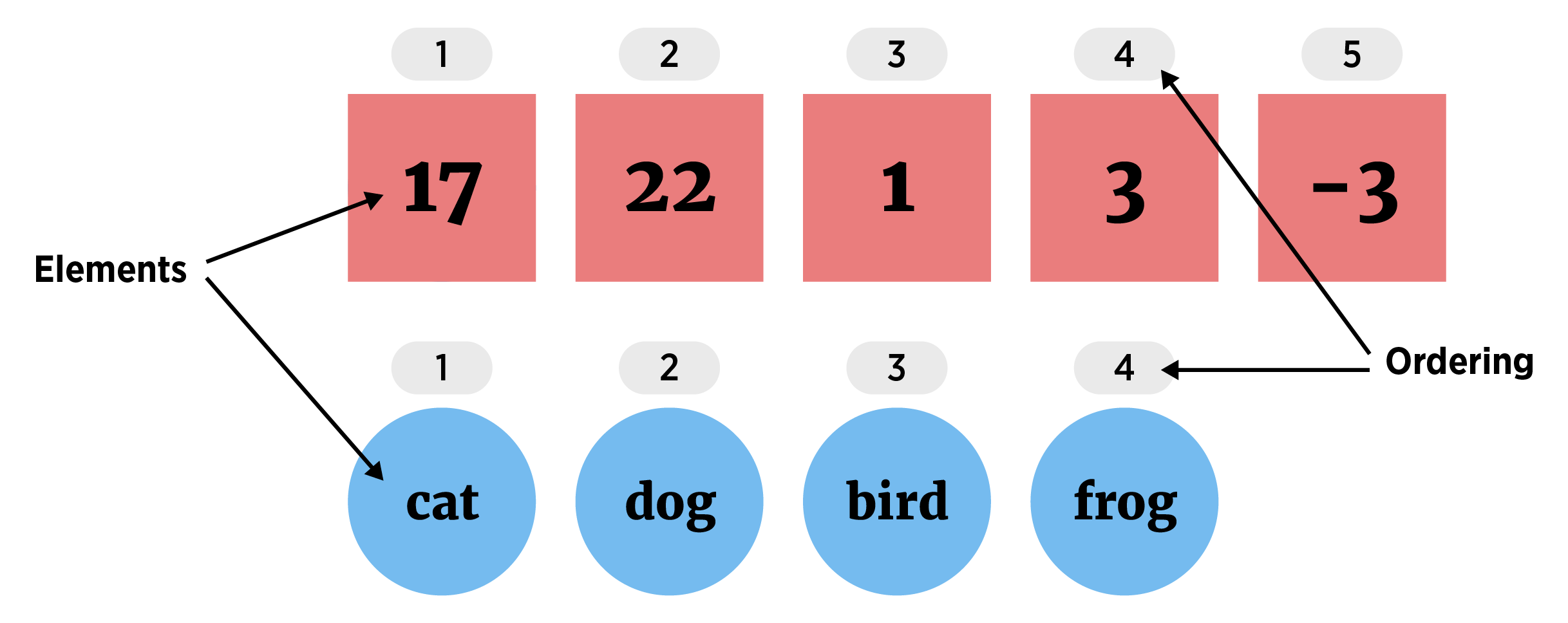
(Atomic) Vector (1D group of elements with an ordering that starts at 1)
Elements must be same the same ‘type’ (homogeneous). The most common types of data are:
- logical, integer, double, and character
Creating a Vector
Many functions output a vector but the most common way to create one yourself is to use the c() function.
c()“combines” values together- Simply separate the values with a comma
#vectors (1 dimensional) objects
#all elements of the same 'type'
x <- c(1, 3, 10, -20, sqrt(2))
x[1] 1.000000 3.000000 10.000000 -20.000000 1.414214- Recall the
str()function to investigate the structure of the object
str(x) num [1:5] 1 3 10 -20 1.41- The
typeof()function tells us which type of data is stored in the vector
typeof(x)[1] "double"- Let’s create another vector
ywith strings stored in it
y <- c("cat", "dog", "bird", "floor")
y[1] "cat" "dog" "bird" "floor"str(y) chr [1:4] "cat" "dog" "bird" "floor"typeof(y)[1] "character"- We can combine two vectors together using
c()as well!
z <- c(x, y)
z[1] "1" "3" "10" "-20"
[5] "1.4142135623731" "cat" "dog" "bird"
[9] "floor" str(z) chr [1:9] "1" "3" "10" "-20" "1.4142135623731" "cat" "dog" "bird" "floor"typeof(z)[1] "character"Notice that R coerces the elements to the data type of the more flexible elements (character - R knows how to convert a number to a character string but doesn’t know how to convert a character string to a number). No warning is produced or message!
You’ll need to get used to how R does these kinds of things implicitly. Check out how R does coercion on some other types of data.
c(TRUE, "hat", 3)[1] "TRUE" "hat" "3" c(c(TRUE, 3), "hat")[1] "1" "3" "hat"Notice that the order of operations above is important to understand! In the first line, R coerces the three elements together (character as it is the most flexible). In the second line, R first coerces TRUE and 3 together. TRUE values are treated as 1 (FALSE values as 0). Then the values of 1 and 3 are coerced to character strings to create the final vector.
- One really useful function that creates a vector is the
seq()(or sequence) function
seq(from = 1, to = 10, by = 2)[1] 1 3 5 7 9seq(1, 10, 1) [1] 1 2 3 4 5 6 7 8 9 10seq(1, 10, length = 5)[1] 1.00 3.25 5.50 7.75 10.00- A shorthand for the
seq()function is to use a:
1:10 [1] 1 2 3 4 5 6 7 8 9 10- This can easily be modified to get other sequences.
20:30/2 [1] 10.0 10.5 11.0 11.5 12.0 12.5 13.0 13.5 14.0 14.5 15.01:15*3 [1] 3 6 9 12 15 18 21 24 27 30 33 36 39 42 45Note that R is doing element-wise math. This is the default behavior of doing math on our common data structures (not just vectors!)
- Another function that creates a vector is the
runif()(random uniform number generator) function.
runif(4, min = 0, max = 1)[1] 0.8019789 0.4866755 0.4202677 0.8751584runif(10) [1] 0.9046116 0.8351989 0.6145934 0.4499420 0.8271097 0.8803008 0.6318299
[8] 0.4316850 0.9439949 0.8358404runif(5, 20, 30)[1] 28.11925 20.97409 29.76169 25.88285 23.51386We’ll find this function really useful when we simulate different quantities!
You might find the different ways to call a function confusing right now! We’ll talk about how to use the help files to understand function calls shortly!
Vector Attributes
R objects can have attributes associated with them. The main attribute that a vector might have associated with it are names for the elements.
u <- c("a" = 1, "b" = 2, "c" = 3)
ua b c
1 2 3 attributes(u)$names
[1] "a" "b" "c"There is a special function for getting at the names of an R object. It is the names() function (nice choice there).
names(u)[1] "a" "b" "c"Names can be useful when it comes to subsetting and matching observations.
Accessing Elements of a Vector
When thinking about accessing (or subsetting) a vector’s elements, remember that vectors are 1D. We can place the numbers corresponding to the positions of the elements we want inside of [] at the end of the vector to return them. Note: R starts its counting at 1, not 0 like many other languages.
- Return vector elements using square brackets
[]at the end of a vector.
letters #built-in vector [1] "a" "b" "c" "d" "e" "f" "g" "h" "i" "j" "k" "l" "m" "n" "o" "p" "q" "r" "s"
[20] "t" "u" "v" "w" "x" "y" "z"letters[1] #R starts counting at 1![1] "a"letters[26][1] "z"- Can ‘feed’ in a vector of indices to
[]
letters[1:4][1] "a" "b" "c" "d"letters[c(5, 10, 15, 20, 25)][1] "e" "j" "o" "t" "y"x <- c(1, 2, 5)
letters[x][1] "a" "b" "e"We’d call x above an indexing vector
- Use negative indices to return a vector without certain elements
letters[-(1:4)] [1] "e" "f" "g" "h" "i" "j" "k" "l" "m" "n" "o" "p" "q" "r" "s" "t" "u" "v" "w"
[20] "x" "y" "z"x <- c(1, 2, 5)
letters[-x] [1] "c" "d" "f" "g" "h" "i" "j" "k" "l" "m" "n" "o" "p" "q" "r" "s" "t" "u" "v"
[20] "w" "x" "y" "z"- If we have a names attribute associated, we can use names to access elements.
ua b c
1 2 3 u["a"]a
1 t <- c("a", "c")
u[t]a c
1 3 Factor Vectors
A factor is a data type used to categorize and store categorical data within a vector. While a standard character string is just a sequence of letters, a factor tells R that the data belongs to a specific, limited set of levels.
For a character string variable where we know the values can only take on a certain set of values, say pet_type: ‘cat’, ‘dog’, ‘bird’, ‘rodent’, or ‘other’, then factors are great! For a variable like name where new values could come in, these aren’t useful.
To create a factor vector we can use factor()!
factor(x = character(), levels, labels = levels,
exclude = NA, ordered = is.ordered(x), nmax = NA)The first argument is the vector of character strings we want to create a vector from. We can then specify the levels or unique values the factor’s elements can take on (and, optionally, different labels for those levels).
pets <- c("cat", "dog", "bird", "cat", "other", "bird", "bird", "other", "dog")
pets_factor <- factor(pets,
levels = c("cat", "dog", "bird", "rodent", "other"))
pets_factor[1] cat dog bird cat other bird bird other dog
Levels: cat dog bird rodent otherstr(pets_factor) Factor w/ 5 levels "cat","dog","bird",..: 1 2 3 1 5 3 3 5 2It may seem odd that the values are actually integers. This comes from the fact that storing integers is easier than storing a character string. We can also compare integers more easily than comparing full character strings.
Notice that we are not able to change the values to something outside of the levels.
pets_factor[1] <- "parrot"Warning in `[<-.factor`(`*tmp*`, 1, value = "parrot"): invalid factor level, NA
generatedThe levels need to correspond to values within the data. We can use the labels argument to have different values show without changing the underlying values.
pets_factor <- factor(pets,
levels = c("cat", "dog", "bird", "rodent", "other"),
labels = c("Cat", "Dog", "Bird", "Rodent", "Others"))
pets_factor[1] Cat Dog Bird Cat Others Bird Bird Others Dog
Levels: Cat Dog Bird Rodent OthersThe other major function to create a factor is as.factor(). This is similar to factor() but doesn’t give the extra arguments (and is sometimes a little faster to run). (In this case, we can’t add the extra level of rodent!)
pets_factor <- as.factor(pets)
pets_factor[1] cat dog bird cat other bird bird other dog
Levels: bird cat dog otherOf course the levels can be modified after the fact!
attributes(pets_factor)$levels
[1] "bird" "cat" "dog" "other"
$class
[1] "factor"There’s a special function to deal with the levels.
levels(pets_factor)[1] "bird" "cat" "dog" "other"We can modify this attribute vector!
levels(pets_factor) <- c(levels(pets_factor), "rodent")
levels(pets_factor)[1] "bird" "cat" "dog" "other" "rodent"(levels() is called a replacement function. These allow you to modify the object by calling it on the left-hand side.)
We’ll come back to factors later on when we look at plotting and summarizing data. The levels can also be ordered which is great when we want to change how a plot or summary looks!
Quick R Video
Let’s look at a quick example of creating and modifying R vectors.
Please pop this video out and watch it in the full panopto player!
Recap!
(Atomic) Vector (1D group of elements with an ordering)
Vectors useful to know about as they are the most basic data object we’ll use
seq()and:Subset vectors using
vec[index]
Use the table of contents on the left or the arrows at the bottom of this page to navigate to the next learning material!